Liver
Optimal platform for drug efficacy and predictive toxicology screening
The pharmaceutical industry is in high need of efficient and relevant in vitro liver models, which can be incorporated in their drug discovery pipelines to identify potential drugs and their toxicity profiles.
Current liver models often rely on cancer cell lines or primary cells, which both have major limitations. The development of human induced pluripotent stem cells (iPSCs) has created a new opportunity for liver disease modelling, drug discovery and liver toxicity research.
DefiniGEN’s highly developed iPSC-derived hepatocyte-like cells, Opti-HEPs, bridge this gap and serve as an optimal platform for drug efficacy and predictive toxicology screening.
DefiniGEN Opti-HEPs have been extensively characterised in response to all the above criteria, revealing their highly metabolic functionality and superiority to existing pre-clinical liver models
DefiniGEN's Opti-HEPs demonstrate characteristic hepatocyte polyhedral, cobblestone-like morphology
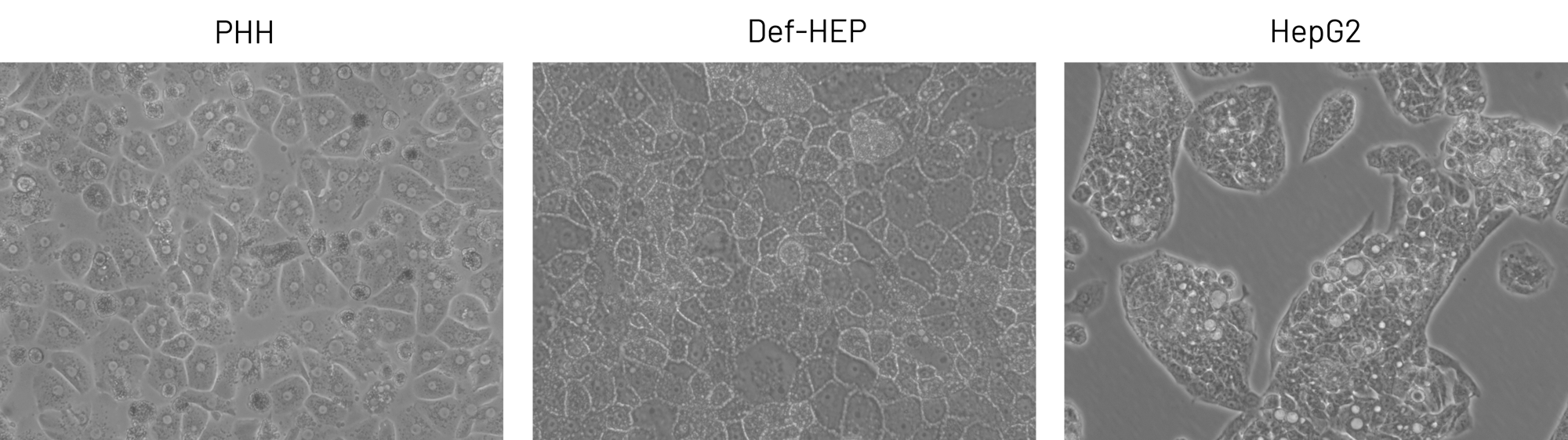
Similar to Primary Human Hepatocytes (PHH), DefiniGEN Opti-HEPs demonstrates the characteristic polyhedral, cobblestone-like morphology and the presence of a uniformed monolayer following >3 weeks of iPSC differentiation, compared to liver carcinoma HepG2 cells. Objective: 20x
DefiniGEN's Opti-HEPs express higher levels of liver maturity markers (albumin, alpha-1-antitrypsin) compared to carcinoma cell lines and are comparable to PHH

A) mRNA expression levels of albumin (ALB) and alpha-1-antitrypsin (A1AT) in liver carcinoma HepG2 cells, Opti-HEP, and primary human hepatocytes (PHH). B) Representative picture of DefiniGEN Opti-HEP revealing protein expression of the hepatocyte maturity markers albumin (red) and alpha-1-antitrypsin (green) by immunocytochemistry (ICC). Nuclei were counterstained with DAPI. Data are presented as mean±SD of n=3-4 independent experiments. mRNA expression data were normalised to PPIA and ICC to total nuclei number. Objective: 10x.
DefiniGEN's Opti-HEPs express higher levels of urea cycle markers and secrete higher levels of urea compared to carcinoma cell lines
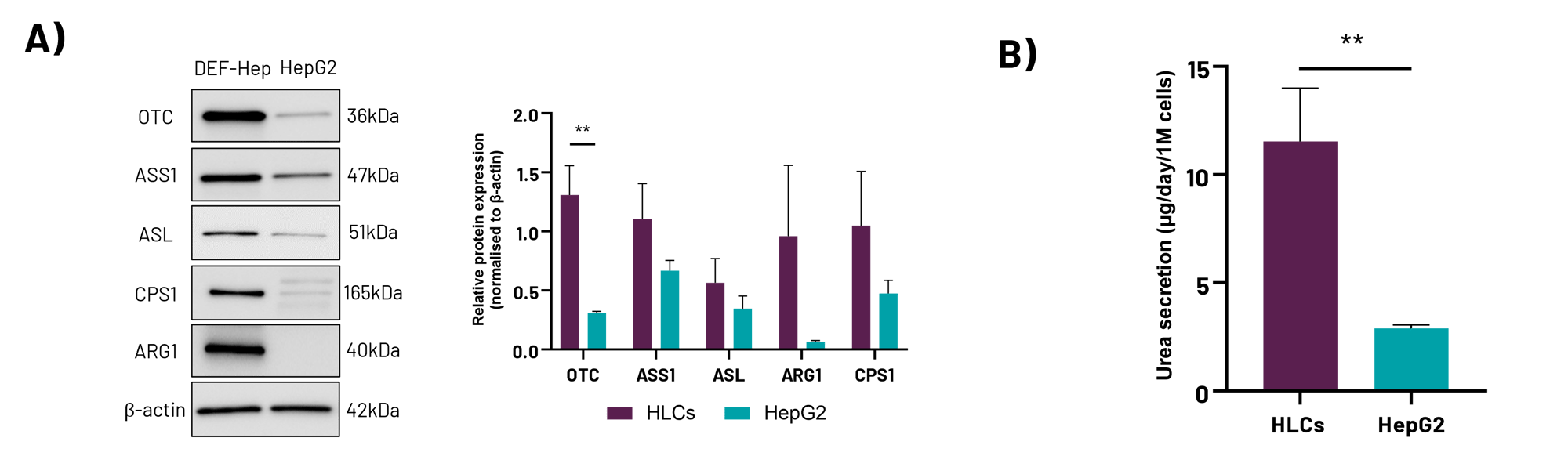
A) Protein expression levels of the urea cycle enzymes OTC, ASS1, ASL, CPS1, and ARG1 in liver carcinoma HepG2 cells and Opti-HEP. B) Urea secretion in Opti-HEPs and liver carcinoma HepG2 cells. Data are presented as mean±SD of n=3-4 independent experiments. Protein expression data were normalised to β- Urea secretion data were normalised to total cell number.
DefiniGEN's Opti-HEPs demonstrate functional carbohydrate and bile acid metabolism pathways
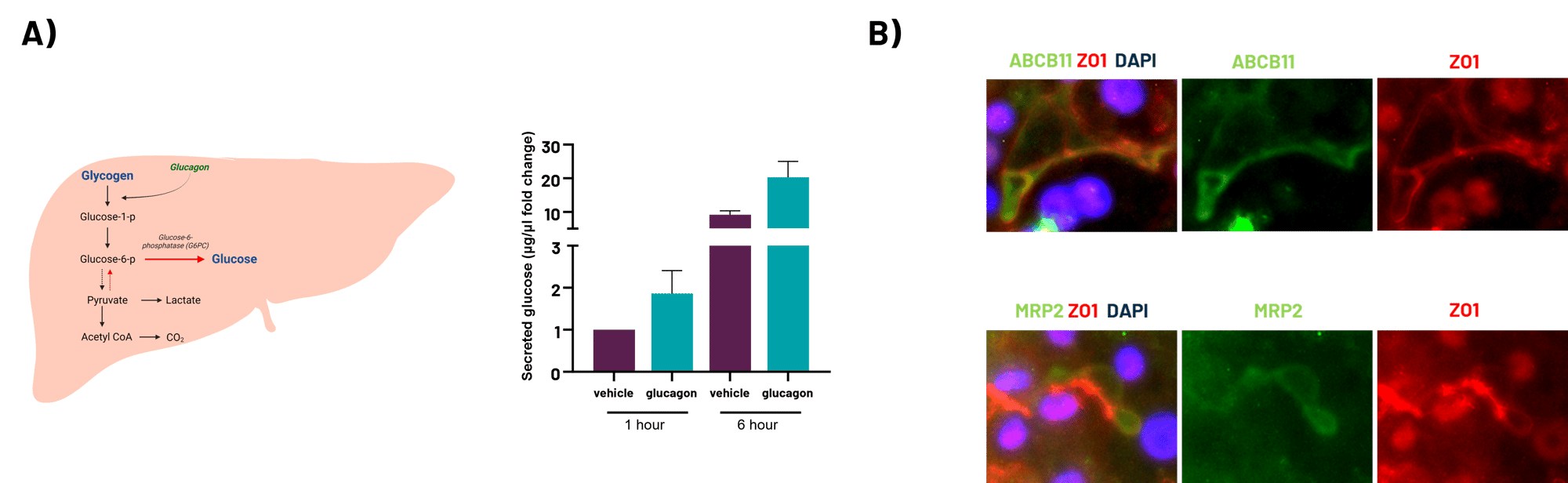
A) Simplified schematic on the gluconeogenesis/glycogenolysis pathways within human liver. Media glucose levels from DefiniGEN Opti-HEP treated with glucose-free media for 1h and 6h with or without glucagon, suggesting the de novo synthesis of glucose from non-lipid precursors, accompanied by functional glycogenolysis pathway additionally contributing to total glucose levels. B) Protein expression of bile acid canaliculi markers ABCB11 and MRP2 (green), co-stained with the apical marker ZO1 (red) in wild-type sandwich cultured Opti-HEP by immunocytochemistry (ICC). Nuclei counterstaining with DAPI (blue). Data are presented as mean±SD of n=2 technical replicates.
DefiniGEN's Opti-HEPs demonstrate cytochrome P450 activity and induction to primary human hepatocytes
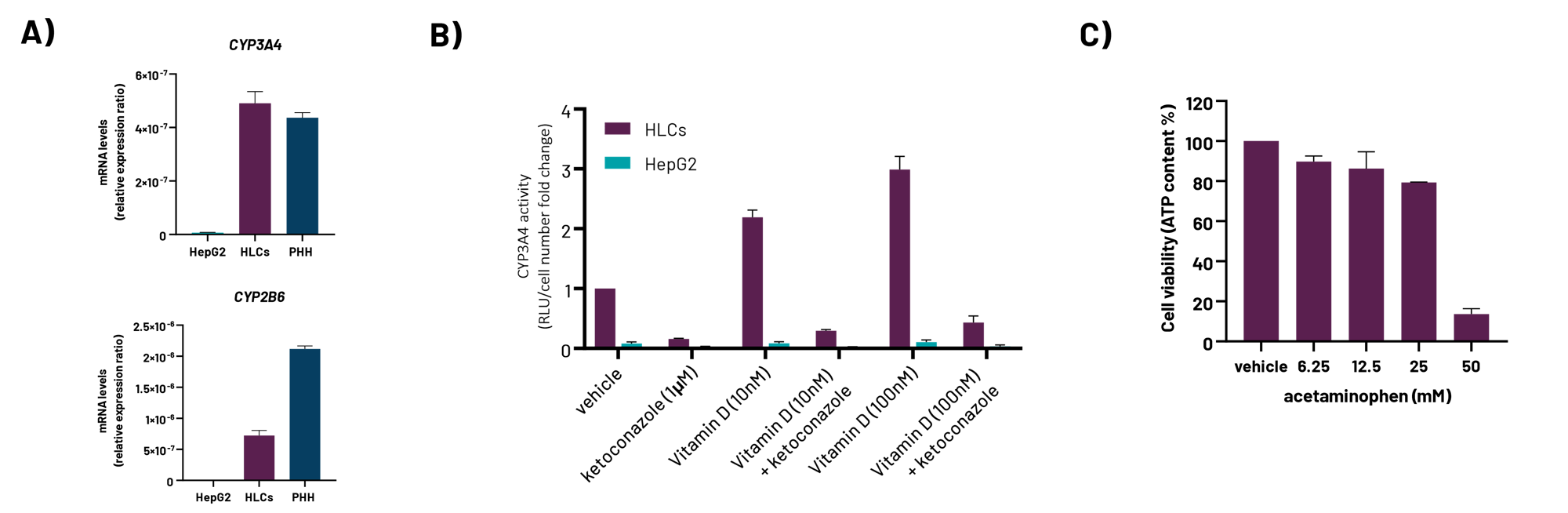
A) mRNA expression levels of CYP3A4 and CYP2B6 in liver carcinoma HepG2 cells, DefiniGEN Opti-HEP, and primary human hepatocytes (PHH).
B) Basal CYP3A4 activity in liver carcinoma HepG2 cells, DefiniGEN Opti-HEP, and PHH.
C) CYP3A4 induction in liver carcinoma HepG2 cells, DefiniGEN Opti-HEP and PHH, following 72h treatment with vehicle, 1μM ketoconazole (CYP3A4 inhibitor), 10-100nM vitamin D (CYP3A4 inducer), or a combination of vitamin D and ketoconazole. mRNA data are presented as mean±SEM of n=2-3 independent experiments and were normalised to 18SrRNA. CYP3A4 activity data are presented as mean±SEM of n=3 independent experiments and were normalised to total cell number.
DefiniGEN's Opti-HEPs demonstrate drug metablism ability
|
No. |
Drug |
FDA labeling |
DILI category, Proctor 2017 |
|
1 |
acetaminophen |
vMost-DILI-Concern |
3. Low Clinical DILI Concern |
|
2 |
amiodarone |
vMost-DILI-Concern |
1. Severe Clinical DILI |
|
3 |
benzbromarone |
vMost-DILI-Concern |
1. Severe Clinical DILI |
|
4 |
bosentan |
vMost-DILI-Concern |
1. Severe Clinical DILI |
|
5 |
carbamazepine |
vMost-DILI-Concern |
2. High Clinical DILI Concern |
|
6 |
clozapine |
vMost-DILI-Concern |
2. High Clinical DILI Concern |
|
7 |
cyclosporin |
vMost-DILI-Concern |
|
|
8 |
diclofenac |
vMost-DILI-Concern |
2. High Clinical DILI Concern |
|
9 |
flutamide |
vMost-DILI-Concern |
1. Severe Clinical DILI |
|
10 |
lapatinib |
vMost-DILI-Concern |
1. Severe Clinical DILI |
|
11 |
nefazodone |
vMost-DILI-Concern |
1. Severe Clinical DILI |
|
12 |
ticrynafen |
vMost-DILI-Concern |
1. Severe Clinical DILI |
|
13 |
troglitazone |
vMost-DILI-Concern |
1. Severe Clinical DILI |
|
14 |
chlorpromazine |
vLess-DILI-Concern |
3. Low Clinical DILI Concern |
|
15 |
desipramine |
Ambiguous DILI-concern |
3. Low Clinical DILI Concern |
|
16 |
fluoxetine |
vLess-DILI-Concern |
3. Low Clinical DILI Concern |
|
17 |
haloperidol |
vLess-DILI-Concern |
|
|
18 |
olanzapine |
vLess-DILI-Concern |
|
|
19 |
paroxetine |
vLess-DILI-Concern |
|
|
20 |
pioglitazone |
vLess-DILI-Concern |
3. Low Clinical DILI Concern |
|
21 |
tacrine |
#N/A |
2. High Clinical DILI Concern |
|
22 |
ambrisentan |
Ambiguous DILI-concern |
5. No DILI |
|
23 |
benztropine |
vNo-DILI-Concern |
5. No DILI |
|
24 |
buspirone |
Ambiguous DILI-concern |
4. Enzyme Elevations in Clinic |
|
25 |
dexamethasone |
Ambiguous DILI-concern |
4. Enzyme Elevations in Clinic |
|
26 |
ethotoin |
Ambiguous DILI-concern |
4. Enzyme Elevations in Clinic |
|
27 |
flavoxate |
vNo-DILI-Concern |
5. No DILI |
|
28 |
hyoscyamine |
#N/A |
5. No DILI |
|
29 |
oxybutynin |
vNo-DILI-Concern |
5. No DILI |
|
30 |
zomepirac |
#N/A |
5. No DILI |
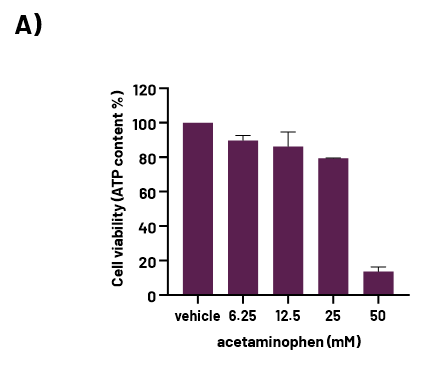
A) Cell viability in Opti-HEP treated with increasing concentrations of acetaminophen (0-50mM).
B) List of compounds ranging from no drug-induced liver toxicity (DILI) to high DILI tested on DefiniGEN Opti-HEP in a dose-dependent manner. Cell viability data are presented as mean±SEM of n=2 and normalised to vehicle.
DefiniGEN’s Opti-HEPs can provide highly predictive in vitro models for efficacy and toxicity studies, minimizing risk while reducing costs, and paving the way for a more efficient and effective future in the field of drug discovery.
Frequently asked questions
Do Opti-HEPs express ASGR1?
Yes, ASGR1 and ASGR2 has been detected by qPCR analysis and ICC and has been shown to localize on the cell membrane.
Do they have a functional urea cycle?
Yes this has been characterized by gene expression profiling, Western blot analysis and Ornithine stimulated functional analysis.
Can you see CYP expression activity?
Expression of CYPs is detectable by qPCR analysis, but they are lower than Primary Human Hepatocytes (PHH).

